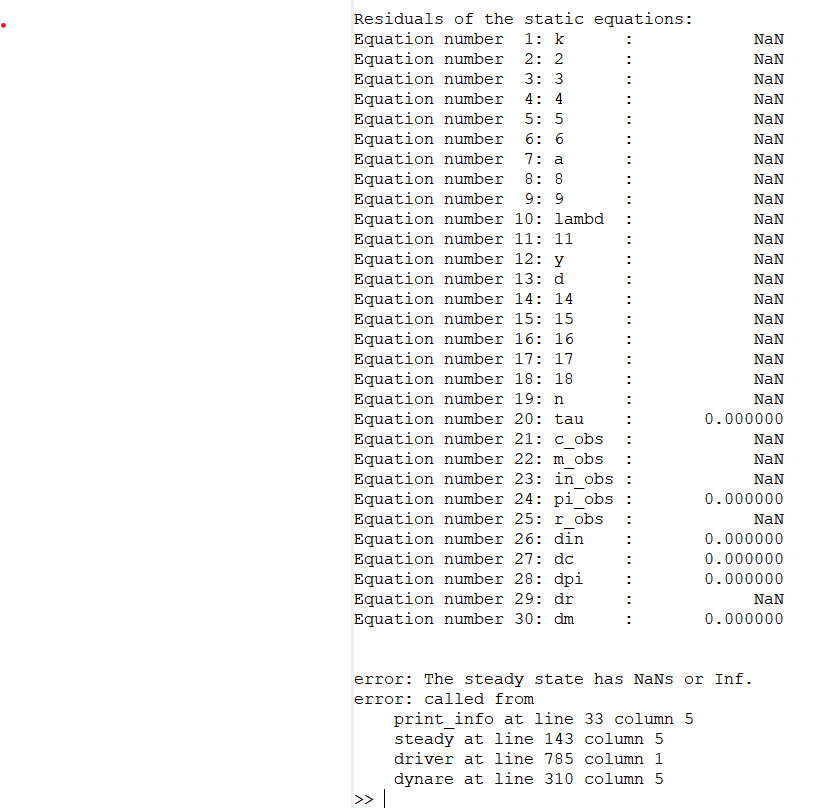
But I was using one-sided filter. Does this mean I can use Kalman filter? My model includes command lik_init = 2 and estimation worked fine at first, till I added parameters sigmaa, sigmae, sigmax, sigmaz and sigmav. Now I obtain error message:
WARNING: Model.mod:1.1-105: Symbol r declared twice.
ERROR: some estimated parameters (sigmaa, sigmae, sigmax, sigmaz, sigmav) also appear in the
expressions defining the variance/covariance matrix of shocks; this is not allowed.
Model.mod (3.8 KB)
Yes, you can use the Kalman filter. lik_init is only about the initialization of the filter. It should not matter. Also, you cannot estimate sigmaa, but rather,
stderr epsilona;
So Kalman filter runs automatically when I add “estimation” line? And it produces full sample maximum likelihood estimates of parameters?
Second, how could I include e and z as parameters, like they are in that paper on page 13/27. Adding them, I obtain error:
ERROR: Model.mod: line 110, col 1 - line 122, col 25: e is not a parameter
Model.mod (3.8 KB)
Yes, at order=1 without OccBin that is the case.
As written above, that makes absolutely no sense.
Perfect, I added option order=1 to the estimation command.
Secondly, I have few questions regarding estimated_params; block. The sigmaa, sigmax, sigmav and sigmaz have inverse gamma distribution according to the paper in the attachment on page 126/262, however stderr epsilona, stderr epsilone and stderr epsilonz have uniform_pdf distribution according to Dynare manual page 20/28. Which one is correct? Moreover, which distribution does beta have (I assume beta distribution?). And omegapi and omegay have normal distribution, does the same go for omegatau? Thank you.
Manual: NTG8.pdf (1.1 MB)
Paper: https://d-nb.info/1026165547/34
Model.mod (3.8 KB)
You can specify pretty much any prior distribution you like.
Thank you. Another question. In my output, which out of four columns are the full sample maximum likelihood parameter estimates, I assume the second column (Mode)?
Zvezek1.xlsx (9.2 KB)
I think you are confusing something. If you specify a (non-uniform) prior, you are not conducting ML, you are conducting Bayesian estimation. But yes, the statistic of interest is the posterior mode.
Of course, this makes perfect sense. But I have to do ML estimation with Kalman filter, according to that paper. Is it possible to do it without major changes in code, by simply specifying uniform distribution for all parameters in estimated_params; block?
As documented in the manual, if you don’t specify any prior, you will be conducting ML estimation.
Thank you, I deleted priors, but I obtain this error message.
Model.mod (3.5 KB)
You did not delete priors:
estimated_params;
bet, beta_pdf, 0.99, 0.01;
clearly specifies a prior distribution. Also, your are calling steady without having initialized the parameter values.
Right, I brought back the section with parameter values in the beginning and deleted the estimated_params; section, but now I obtain error:
Estimation: the ‘estimated_params’ block is mandatory (unless you are running a smoother)
Model.mod (3.2 KB)
Dear professor @jpfeifer or @sebastien,
can you please give me advice on how to run maximum likelihood estimation. I deleted priors (section estimated_params) but now I obtain error message:
Estimation: the ‘estimated_params’ block is mandatory (unless you are running a smoother)
I checked the manual on page 71/223, but this information isn’t there.
Thank you very much in advance!
Model.mod (3.2 KB)
The block is mandatory, but if no statistical PDF is specified, MLE is implied because there is no defined prior distribution.
So the right way is to include empty block like:
estimated_params;
end;
Or should I include also parameters, but without PDF, like
bet, 0.99, 0.01;
instead of
bet, beta_pdf, 0.99, 0.01;
Model.mod (3.2 KB)
Perfect, thank you very much. However, now I obtain an error:
ERROR: Model.mod: line 110, col 16: syntax error, unexpected ‘;’, expecting COMMA
Model.mod (3.6 KB)
Not sure I understand what you are trying to achieve here. This block is documented here:
https://www.dynare.org/manual/the-model-file.html#estimated_params
In a ML context, each estimated parameter must be followed by 1 scalar (the initial guess for the maximisation of the likelihood) or three (comma separated) scalars (the initial guess, a lower bound and an upper bound defining the set of possible values for the parameters if you want to restrict the ML estimator). In your case you have only two numbers after beta, this is not allowed.
Best,
Stéphane
Thank you, my code works now!