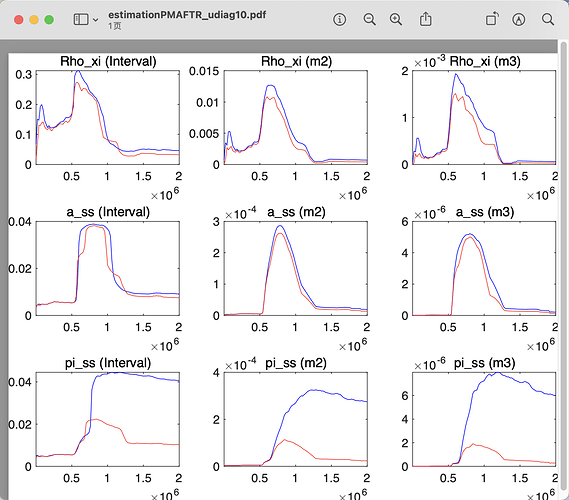
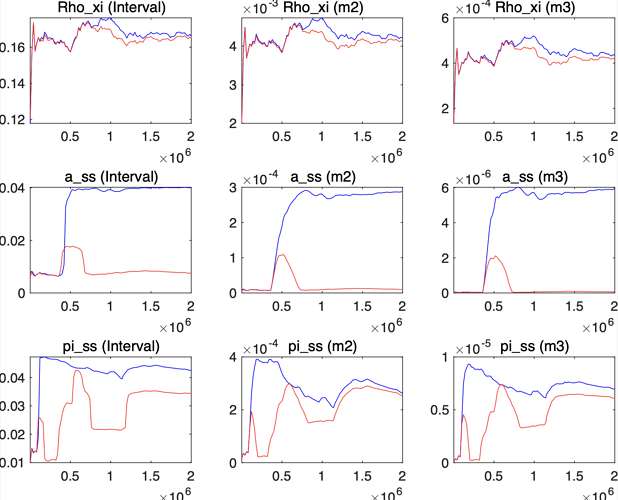
I get such result in convergence diagnostic, it seems never get to convergence even give bigger mh_replic.
there are several question follow:
- Why do this happen, where do I need to check? Does this means there’s something I do wrong?
- How can I improve this? (Use same data different model the convergence diagnostic look good)
regards
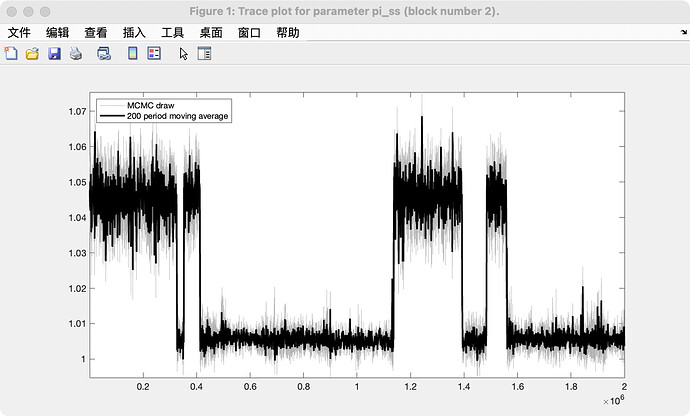
Did you have a look at the trace_plots?
for pi_ss here, trace_plot:
Your model shows bimodality. That explains the convergence issues.
Thanks for reply. I am still confused.
Do the two curve of diagnostic must converge to be consistent?
How may I improve that?
Do this means I have done something wrong?
regards!
In principle, yes. But with two chains and two modes, it will be almost impossible to get the desired result due to the switches between modes that are also random. I would rather go for a vary long chain.
Thank you sir.
If I use 2 different model to do estimation, and one of the diagnostic looks good(Model A) while the other fails like above(Model B), then the good-looking one would be a better choice. And the bimodality is from the switches between different modes(same model but in different parameters value). Is this understanding correct?
And in same situation, do the log-data-density of the model in bimodality still precise? If I am comparing these two, and the A’s log data density > B’s, can I give a conclusion that comparing to B, A suits the data more?
Thanks in advance!
Hope I haven’t take up too much of your time
Evaluation of the model fit requires correctly sampling from the posterior. If that does not work yet, you cannot draw any conclusions. The marginal data density can be used to compare models. The Laplace approximation relies on one mode, so it will be wrong. The modified harmonic mean should work, but only if you are correctly sampling from the posterior.
Thanks sir, please forgive me to ask this: How would I check if I am correctly sampling from the posterior?
That is the tricky part, given your bimodality. That’s why I suggested a really long chain instead of two chains.